Working with unyt¶
Basic Usage¶
To use unyt in a project:
>>> import unyt
The top-level unyt namespace defines both a number of useful functions as
well as a number of units and physical constants populated from the
unyt.unit_symbols and unyt.physical_constants namespaces you can
use to attach units to NumPy arrays and other common Python data container types
like list and tuple. For an exhaustive listing of units and physical
constants defined in unyt, see Listing of Units.
Warning
Both unit symbols and physical constants are defined in the top-level
unyt namespace. Some names occur as both unit symbols and physical
constants, e.g. "me", "mp", "E_pl", etc. In such cases, the
top-level namespace defaults to exporting the physical constant with the
duplicate name. This means that there may be a (very rare) case where an
Unit object that at one point can be
imported from the top-level namespace may at a later point be only imported
as an unyt_quantity object with the
same name. As described below, there are other ways to import the unit
symbols and/or physical constants besides the top-level namespace.
An Example from High School Physics¶
To see how you might use unyt to solve a problem where units might be a
headache, let’s estimate the orbital periods of Jupiter’s Galilean moons,
assuming they have circular orbits and their masses are negligible compared to
Jupiter. Under these assumptions, the orbital period is
For this exercise let’s calculate the orbital period in days. While it’s
possible to do this using plain old floating point numbers (you probably had to
do something similar on a calculator in a high school physics class, looking up
and plugging in conversion factors by hand), it’s much easier to do this sort of
thing symbolically and let unyt handle the unit conversions.
To do this we’ll need to know the mass of Jupiter (fortunately that is built
into unyt) and the semi-major axis of the orbits of Jupiter’s moons, which
we can look up from Wikipedia and enter by hand:
>>> from unyt import Mjup, G, AU
>>> from math import pi
...
>>> moons = ['Io', 'Europa', 'Ganymede', 'Callisto']
>>> semimajor_axis = [.002819, .0044856, .00715526, .01258513]*AU
...
>>> period = 2*pi*(semimajor_axis**3/(G*Mjup))**0.5
>>> period = period.to('d')
...
>>> for moon, period in zip(moons, period):
... print('{}: {:04.2f}'.format(moon, period))
Io: 1.77 day
Europa: 3.55 day
Ganymede: 7.15 day
Callisto: 16.69 day
Let’s break up this example into a few components so you can see what’s going
on. First, we import the unit symbols we need from the unyt namespace:
>>> from unyt import Mjup, G, km
The unyt namespace has a large number of units and physical constants you
can import to apply units to data in your own code. You can see how that works
in the example:
>>> semimajor_axis = [.002819, .0044856, .00715526, .01258513]*AU
>>> semimajor_axis
unyt_array([0.002819 , 0.0044856 , 0.00715526, 0.01258513], 'AU')
By multiplying by km, we converted the Python list into a
unyt.unyt_array instance. This is a class
that’s built into unyt, has units attached to it, and knows how to
convert itself into different dimensionally equivalent units:
>>> semimajor_axis.value
array([0.002819 , 0.0044856 , 0.00715526, 0.01258513])
>>> semimajor_axis.units
AU
>>> print(semimajor_axis.to('km'))
[ 421716.39764641 671036.20903964 1070411.66066813 1882708.6511216 ] km
Next, we calculated the orbital period by translating the orbital period formula to Python and then converting the answer to the units we want in the end, days:
>>> period = 2*pi*(semimajor_axis**3/(G*Mjup))**0.5
>>> period
unyt_array([ 152864.59689789, 306828.08975058, 618162.17963649,
1441952.18891597], 's')
>>> period.to('d')
unyt_array([ 1.76926617, 3.55125104, 7.15465486, 16.68926145], 'day')
Note that we haven’t added any conversion factors between different units,
that’s all handled internally by unyt. Also note how the
unyt_array.to method was able to
automatically handle the conversion from seconds to days and how the
shorthand "d" was automatically interpreted as "day".
Arithmetic and units¶
The real power of working with unyt is its ability to add, subtract,
multiply, and divide quantities and arrays with units in mathematical formulas
while automatically handling unit conversions and detecting when you have made a
mistake in your units in a mathematical formula. To see what I mean by that,
let’s take a look at the following examples:
>>> from unyt import cm, m, ft, yard
>>> print(3.*cm + 4.*m - 5.*ft + 6.*yard)
799.24 cm
Despite the fact that the four unit symbols used in the above example correspond
to four different units, unyt is able to automatically convert the value
of all three units into a common unit and return the result in those units. Note
that for expressions where the return units are ambiguous, unyt always
returns data in the units of the leftmost object in an expression:
>>> print(4*m + 3*cm - 5*ft + 6*yard)
7.9924 m
One can also form more complex units out of atomic unit symbols. For example, here is how we’d create an array with units of meters per second and print out the values in the array in miles per hour:
>>> from unyt import m, s
>>> velocities = [20., 22., 25.]*m/s
>>> print(velocities.to('mile/hr'))
[44.73872584 49.21259843 55.9234073 ] mile/hr
Similarly one can multiply two units together to create new compound units:
>>> from unyt import N, m
>>> energy = 3*N * 4*m
>>> print(energy)
12 N*m
>>> print(energy.to('erg'))
120000000.0 erg
In general, one can multiply or divide by an arbitrary rational power of a unit symbol. Most commonly this shows up in mathematical formulas in terms of square roots. For example, let’s calculate the gravitational free-fall time for a person to fall from the surface of the Earth through to a hole dug all the way to the center of the Earth. It turns out that this time is given by:
where \(\rho\) is the average density of the Earth.
>>> from unyt import G, Mearth, Rearth
>>> from math import pi
>>> import numpy as np
...
>>> rho = Mearth / (4./3 * pi* Rearth**3)
>>> print(rho.to('g/cm**3'))
5.581225129861083 g/cm**3
>>> tff = np.sqrt(3*pi/(32*G*rho))
>>> print(tff.to('min'))
14.820022043294829 min
If you make a mistake by adding two things that have different dimensions,
unyt will raise an error to let you know that you have a bug in your
code:
>>> from unyt import kg, m
>>> 3*kg + 5*m
Traceback (most recent call last):
...
unyt.exceptions.UnitOperationError: The <ufunc 'add'> operator for
unyt_arrays with units "kg" (dimensions "(mass)") and
"m" (dimensions "(length)") is not well defined.
while this example is trivial when one writes more complicated formulae it can be easy to accidentally write expressions that are not dimensionally sound.
Sometimes this can be annoying to deal with, particularly if one is mixing data
that has units attached with data from some outside source with no units. To
quickly patch over this lack of unit metadata (which could be applied by
explicitly attaching units at I/O time), one can use the units attribute of
the unyt.unyt_array class to quickly apply units to a scalar, list, or array:
>>> from unyt import cm, s
>>> velocities = [10, 20, 30] * cm/s
>>> velocities + 12
Traceback (most recent call last):
...
unyt.exceptions.UnitOperationError: The <ufunc 'add'> operator for
unyt_arrays with units "cm/s" (dimensions "(length)/(time)") and
"dimensionless" (dimensions "1") is not well defined.
>>> velocities + 12*velocities.units
unyt_array([22, 32, 42], 'cm/s')
Powers, Logarithms, Exponentials, and Trigonometric Functions¶
The unyt library represents powers using standard Python syntax. This
means you must use ** and not ^, even when writing a unit as a string:
>>> from unyt import kg, m
>>> print((10.*kg/m**3).to('g/cm**3'))
0.01 g/cm**3
Formally it does not make sense to exponentiate, take the logarithm of, or apply
a transcendental function to a quantity with units. However, the unyt
library makes the practical affordance to allow this, simply ignoring the units
present and returning a result without units. This makes it easy to work with
data that has units both in linear space and in log space:
>>> from unyt import g, cm
>>> import numpy as np
>>> print(np.log10(1e-23*g/cm**3))
-23.0
The one exception to this rule is for trigonometric functions applied to data with angular units:
>>> from unyt import degree, radian
>>> import numpy as np
>>> np.sin(np.pi/4*radian)
array(0.70710678)
>>> np.sin(45.*degree)
array(0.70710678)
Logarithmic Quantities and Units¶
The logarithmic quantities level-of-power and level-of-field and the units neper and
bel are supported. In the next example, we represent the power measurements, p, as
a logarithmic quantity at reference level, p_ref, in the units decibel.
>>> import numpy as np
>>> from unyt import dB, mW
>>> dB.dimensions
(logarithmic)
>>> p = [1, 100]*mW
>>> p_ref = 1*mW
>>> level_of_power = 10*np.log10(p/p_ref)*dB
>>> level_of_power
unyt_array([ 0., 20.], 'dB')
You can convert the logarithmic quantity back to physical units through exponentiation,
just remember to remove the units using the
unyt_array.v property.
>>> 10**(level_of_power.v/10)*p_ref
unyt_array([ 1., 100.], 'mW')
Printing Units¶
The print formatting of unyt_array can be
controlled identically to NumPy arrays, using numpy.setprintoptions:
>>> import numpy as np
>>> import unyt as u
...
>>> np.set_printoptions(precision=4)
>>> print([1.123456789]*u.km)
[1.1235] km
>>> np.set_printoptions(precision=8)
Print a \(\rm{\LaTeX}\) representation of a set of units using the
unyt.unit_object.Unit.latex_representation() function or
unyt.unit_object.Unit.latex_repr attribute:
>>> from unyt import g, cm
>>> (g/cm**3).units.latex_representation()
'\\frac{\\rm{g}}{\\rm{cm}^{3}}'
>>> (g/cm**3).units.latex_repr
'\\frac{\\rm{g}}{\\rm{cm}^{3}}'
Simplifying Units¶
Unit expressions can often be simplified to cancel pairs of factors with compatible dimensions. For example, we can form a unit with dimensions of length by dividing a unit with dimensions of length squared by another unit with dimensions of length:
>>> from unyt import m, cm
>>> m**2/cm
m**2/cm
The Unit class has a simplify() method that we can call to create a new unit
object to that includes the dimensionless ratio m/cm as a constant
coefficient:
>>> (m**2/cm).simplify()
100*m
This will also work for units that are the reciprocals of each other, for example:
>>> from unyt import s, Hz
>>> (s*Hz).simplify()
(dimensionless)
Products and quotients of unit objects will not be simplified unless
simplify() is called explicitly. However, products and quotients of arrays
and quantities will be simplified to make interactive work more intuitive:
>>> from unyt import erg, minute, hour
>>> power = [20, 40, 80] * erg / minute
>>> elapsed_time = 3*hour
>>> print(power*elapsed_time)
[ 3600. 7200. 14400.] erg
Checking Units¶
If you write a function that accepts data with units as an argument or returns data with units,
you can ensure the dimensional correctness of the inputs or outputs
using the @accepts and @returns decorators:
>>> from unyt.dimensions import length, time
>>> from unyt import accepts, returns
>>> import unyt as u
>>> @returns(length)
... @accepts(a=time, v=length/time)
... def foo(a, v):
... return a * v
...
>>> res = foo(a=2*u.s, v=3*u.m/u.s)
>>> print(res)
6 m
@accepts can specify the dimensions of any subset of inputs and @returns must always describe all outputs.
>>> @returns(length, length/time**2)
... @accepts(v=length/time)
... def bar(a, v):
... return a * v, v / a
...
>>> res = bar(a=2*u.s, v=3*u.m/u.s)
>>> print(*res)
6 m 1.5 m/s**2
Note
Using these decorators may incur some performance overhead, especially for small arrays.
Temperature Units¶
The temperature unit degree Celsius has the symbol °C, but since the degree character
is an invalid Python identifier, unyt uses the symbol degC. Printing a degree Celsius
quantity will show the correct symbol.
>>> from unyt import degC
>>> Ta = 23*degC
>>> print(Ta)
23 °C
The degC symbol has alternative names degree_Celsius, Celsius and °C.
>>> from unyt import degree_Celsius, unyt_array
>>> Ta = 23*degree_Celsius
>>> print(Ta)
23 °C
>>> Ta = unyt_array([-40, 23, 70], '°C')
>>> print(Ta)
[-40 23 70] °C
These comments also apply to degree Fahrenheit.
Performing arithmetic with temperature quantities can be ambiguous. To clarify intent,
unyt has the convenience units delta_degC and delta_degF.
>>> from unyt import degC, delta_degC, V
>>> t1 = 23*degC
>>> t2 = 1*delta_degC
>>> print(t1 + t2)
24.0 °C
>>> print(t2 - t1)
-22.0 °C
>>> tempco = 10.0*V/delta_degC
>>> print(tempco*2*delta_degC)
20.0 V
Unit Conversions and Unit Systems¶
Converting Data to Arbitrary Units¶
If you have some data that you want to convert to a different set of units and
you know which units you would like to convert it to, you can make use of the
unyt_array.to function:
>>> from unyt import mile
>>> (1.0*mile).to('ft')
unyt_quantity(5280., 'ft')
If you try to convert to a unit with different dimensions, unyt will
raise an error:
>>> from unyt import mile
>>> (1.0*mile).to('lb')
Traceback (most recent call last):
...
unyt.exceptions.UnitConversionError: Cannot convert between 'mile' (dim
'(length)') and 'lb' (dim '(mass)').
While we recommend using unyt_array.to in
most cases to convert arrays or quantities to different units, if you would like
to explicitly emphasize that this operation has to do with units, we also
provide the more verbose name unyt_array.in_units which behaves identically to
unyt_array.to.
Converting Units In-Place¶
The unyt_array.to method makes a copy of the
array data. For most cases this is fine, but when dealing with big arrays, or
when performance is a concern, it sometimes is preferable to convert the data in
an array in-place, without copying the data to a new array. This can be
accomplished with the unyt_array.convert_to_units function:
>>> from unyt import mile
>>> data = [1., 2., 3.]*mile
>>> data
unyt_array([1., 2., 3.], 'mile')
>>> data.convert_to_units('km')
>>> data
unyt_array([1.609344, 3.218688, 4.828032], 'km')
Converting to MKS and CGS Base Units¶
If you don’t necessarily know the units you want to convert data to ahead of
time, it’s often convenient to specify a unit system to convert to. The
unyt_array has built-in conversion methods for
the two most popular unit systems, MKS (meter kilogram second) and CGS
(centimeter gram second). For CGS these are unyt_array.in_cgs and unyt_array.convert_to_cgs. These functions create a new copy of an
array in CGS units and convert an array in-place to CGS respectively. For MKS,
there are the unyt_array.in_mks
and unyt_array.convert_to_mks methods, which play analogous roles.
See below for details on CGS and MKS electromagnetic units.
Metallicity Unit Conversions¶
In the astrophysical context, “metals” are all of the elements that have atomic
numbers greater than 2, i.e. everything heavier than helium. The “solar metallicity”
is the mass fraction of metals in the solar atmosphere, and is used in a variety
of contexts. Often, the metallicity of other astrophysical objects is expressed
in terms of the solar metallicity, given by the unit \(Z_\odot\). The default
mass fraction corresponding to \(Z_\odot\) in unyt is 0.01295, corresponding
to the value used in the Cloudy Code.
Metal mass fractions (by definition dimensionless) can be converted to \(Z_\odot\)
(and vice versa):
>>> from unyt import dimensionless
>>> M_Z = 0.0259*dimensionless
>>> M_Z
unyt_quantity(0.0259, 'dimensionless')
>>> M_Z.convert_to_units("Z_sun")
>>> M_Z
unyt_quantity(2., 'Zsun')
However, the value of this mass fraction conversion must be measured, and various
estimates of it disagree somewhat. Different sub-disciplines of astronomy often
use different estimates in the literature. unyt provides other metallicity
unit conversions to several typical values in use. The available units (and their
mass fraction conversion factors) are:
"Zsun_angr": 0.01937, from Anders E. & Grevesse N. (1989, Geochimica et Cosmochimica Acta 53, 197)"Zsun_aspl": 0.01337, from Asplund M., Grevesse N., Sauval A.J. & Scott P. (2009, ARAA, 47, 481)"Zsun_feld": 0.01909, from Feldman U. (1992, Physica Scripta, 46, 202)"Zsun_lodd": 0.01321, from Lodders, K (2003, ApJ 591, 1220)
These can be used in the same way as above:
>>> from unyt import dimensionless
>>> M_Z = 0.0259*dimensionless
>>> M_Z
unyt_quantity(0.0259, 'dimensionless')
>>> M_Z.convert_to_units("Zsun_angr")
>>> M_Z
unyt_quantity(1.33711926, 'Zsun_angr')
Other Unit Systems¶
The unyt library currently has built-in support for a number of unit
systems, as detailed in the table below. Note that all unit systems currently
use “radian” as the base angle unit.
If a unit system in the table below has “Other Units” specified, this is a mapping from dimension to a unit name. These units override the unit system’s default unit for that dimension. If no unit is explicitly specified of a dimension then the base unit for that dimension is calculated at runtime by combining the base units for the unit system into the appropriate dimension.
Unit system |
Base Units |
Other Units |
|---|---|---|
cgs |
cm, g, s |
|
mks |
m, kg, s |
|
imperial |
ft, lb, s |
|
galactic |
kpc, Msun, kyr |
|
solar |
AU, Mearth, yr |
Note that in MKS units the current unit, Ampere, is a base unit in the unit
system. In CGS units the electromagnetic units like Gauss and statA are
decomposable in terms of the base mass, length, and time units in the unit
system. For this reason quantities defined in E&M units in CGS units are not
readily convertible to MKS units and vice versa since the units are not
dimensionally equivalent. The unyt library does have limited support for converting electromagnetic units between MKS and CGS, however only simple conversions of data with a single specific unit are supported and no conversions are allowed for complex combinations of units. For example converting between Gauss and Tesla is supported:
>>> from unyt import T
>>> (1.0*T).to('G')
unyt_quantity(10000., 'G')
But converting a more complicated compound unit will raise an error:
>>> from unyt import C, T, V
>>> (1.0*C*T*V).in_cgs()
Traceback (most recent call last):
...
unyt.exceptions.UnitsNotReducible: The unit "C*T*V" (dimensions
"(length)**2*(mass)**2/((current_mks)*(time)**4)") cannot be reduced to
an expression within the cgs system of units.
If you need to work with complex expressions involving electromagnetic units, we
suggest sticking to either CGS or SI units for the full calculation. There is no
general way to convert an arbitrary quantity between CGS and SI units if the
quantity involves electromagnetic units. Instead, it is necessary to do the
conversion on the equations under consideration, and then recompute the
necessary quantity in the transformed set of equations. This requires
understanding the context for a calculation, which unfortunately is beyond the
scope of a library like unyt.
You can convert data to a unit system unyt knows about using the
unyt_array.in_base and
unyt_array.convert_to_base
methods:
>>> from unyt import g, cm, horsepower
>>> (1e-9*g/cm**2).in_base('galactic')
unyt_quantity(4.78843804, 'Msun/kpc**2')
>>> data = [100., 500., 700.]*horsepower
>>> data
unyt_array([100., 500., 700.], 'hp')
>>> data.convert_to_base('mks')
>>> data
unyt_array([ 74569.98715823, 372849.93579114, 521989.91010759], 'W')
Defining and Using New Unit Systems¶
To define a new custom unit system, one need only create a new instance of the
unyt.UnitSystem class. The class
initializer accepts a set of base units to define the unit system. If you would
like to additionally customize any derived units in the unit system, you can do
this using item setting.
As an example, let’s define an atomic unit system based on typical scales for atoms and molecules:
>>> from unyt import UnitSystem
>>> atomic_unit_system = UnitSystem('atomic', 'nm', 'mp', 'fs', 'nK', 'rad')
>>> atomic_unit_system['energy'] = 'eV'
>>> atomic_unit_system
atomic Unit System
Base Units:
length: nm
mass: mp
time: fs
temperature: nK
angle: rad
current_mks: A
luminous_intensity: cd
logarithmic: Np
Other Units:
energy: eV
>>> print(atomic_unit_system)
atomic
>>> atomic_unit_system['number_density']
nm**(-3)
>>> atomic_unit_system['angular_momentum']
mp*nm**2/fs
It is also legal to define a unit system using unyt.Unit instances:
>>> from unyt.unit_symbols import Msun, second, megaparsec
>>> UnitSystem('cosmological', megaparsec, Msun, second)
cosmological Unit System
Base Units:
length: Mpc
mass: Msun
time: s
temperature: K
angle: rad
current_mks: A
luminous_intensity: cd
logarithmic: Np
Other Units:
Or with a quantity:
>>> UnitSystem('quasmological', 3*megaparsec, .8*Msun, 42*second)
quasmological Unit System
Base Units:
length: 3*Mpc
mass: 0.8*Msun
time: 42*s
temperature: K
angle: rad
current_mks: A
luminous_intensity: cd
logarithmic: Np
Other Units:
Once you have defined a new unit system that will register the new system with a
global registry of unit systems known to the unyt library. That means you
will immediately be able to use it just like the built-in unit systems:
>>> from unyt import W
>>> (1.0*W).in_base('atomic')
unyt_quantity(0.59746607, 'mp*nm**2/fs**3')
If you would like your unit system to include an MKS current unit
(e.g. something that is directly convertible to the MKS Ampere unit), then
specify a current_mks_unit in the UnitSystem initializer.
Equivalencies¶
An equivalency is a way to define a mapping to convert from one unit to another even if the two units are not dimensionally equivalent. This usually involves some sort of shorthand or heuristic understanding of the problem under consideration. Only use one of these equivalencies if it makes sense to use it for the problem you are working on.
The unyt library implements the following equivalencies:
"thermal": conversions between temperature and energy (\(E = k_BT\))"spectral": conversions between wavelength, spatial frequency, frequency, and energy for photons (\(E = h\nu = hc/\lambda\), \(c = \lambda\nu\))"mass_energy": conversions between mass and energy (\(E = mc^2\))"lorentz": conversions between velocity and Lorentz factor (\(\gamma = 1/\sqrt{1-(v/c)^2}\))"schwarzschild": conversions between mass and Schwarzschild radius (\(R_S = 2GM/c^2\))"compton": conversions between mass and Compton wavelength (\(\lambda = h/mc\))
You can convert data to a specific set of units via an equivalency appropriate
for the units of the data. To see the equivalencies that are available for an
array, use the unit_array.list_equivalencies method:
>>> from unyt import gram, km
>>> gram.list_equivalencies()
mass_energy: mass <-> energy
schwarzschild: mass <-> length
compton: mass <-> length
>>> km.list_equivalencies()
spectral: length <-> spatial_frequency <-> frequency <-> energy
schwarzschild: mass <-> length
compton: mass <-> length
All of the unit conversion methods described above have an equivalence
keyword argument that allows one to optionally specify an equivalence to use for
the unit conversion operation. For example, let’s use the schwarzschild
equivalence to calculate the mass of a black hole with a radius of one AU:
>>> from unyt import AU
>>> (1.0*AU).to('Msun', equivalence='schwarzschild')
unyt_quantity(50656851.7815179, 'Msun')
Both the methods that convert data in-place and the ones that return a copy
support optionally specifying equivalence. In addition to the methods described
above, unyt also supplies two more conversion methods that require an
equivalence to be specified: unyt_array.to_equivalent and
unyt_array.convert_to_equivalent. These are identical to their
counterparts described above, except that equivalence is a required
argument to the function rather than an optional keyword argument. Use these
functions when you want to emphasize that an equivalence is being used.
If the equivalence has optional keyword arguments, these can be passed to the
unit conversion function. For example, here’s an example where we specify a
custom mean molecular weight (mu) for the number_density equivalence:
>>> from unyt import g, cm
>>> rho = 1e-23 * g/cm**3
>>> rho.to('cm**-3', equivalence='number_density', mu=1.4)
unyt_quantity(4.26761476, 'cm**(-3)')
For full API documentation and an autogenerated listing of the built-in
equivalencies in unyt as well as a short usage example for each, see the
unyt.equivalencies API listing.
Dealing with code that doesn’t use unyt¶
Optimally, a function will work the same irrespective of whether the data passed in has units attached or not:
>>> from unyt import cm
>>> def square(x):
... return x**2
>>> print(square(3.))
9.0
>>> print(square(3.*cm))
9.0 cm**2
However in the real world that is not always the case. In this section we describe strategies for dealing with that situation.
Stripping units off of data¶
The unyt library provides a number of ways to convert
unyt_quantity instances into floats and
unyt_array instances into NumPy arrays. These
methods either return a copy of the data as a NumPy array or return a view
onto the underlying array data owned by a unyt_array instance.
To obtain a new array containing a copy of the original data, use either the
unyt_array.to_value function or the
unyt_array.value or unyt_array.v properties. All of these are equivalent to passing a
unyt_array to the numpy.array() function:
>>> from unyt import g
>>> import numpy as np
>>> data = [1., 2., 3.]*g
>>> data
unyt_array([1., 2., 3.], 'g')
>>> np.array(data)
array([1., 2., 3.])
>>> data.to_value('kg')
array([0.001, 0.002, 0.003])
>>> data.value
array([1., 2., 3.])
>>> data.v
array([1., 2., 3.])
Similarly, to obtain a ndarray containing a view of the data in the original
array, use either the unyt_array.ndview
property (or unyt_array.d for shorts):
>>> data.view(np.ndarray)
array([1., 2., 3.])
>>> data.ndview
array([1., 2., 3.])
>>> data.d
array([1., 2., 3.])
Applying units to data¶
Note
A NumPy array that shares memory with another NumPy array points to the array
that owns the data with the base attribute. If arr1.base is arr2 is
True then arr1 is a view onto arr2 and arr2.base will be
None.
When a unyt_array instance is created from a
NumPy array and a Unit, data from the NumPy array
will be copied:
>>> from unyt import g
>>> data = np.random.random((100, 100))
>>> data_with_units = data*g
>>> data_with_units.base is data
False
If you would like to create a view rather than a copy, you can apply units like this:
>>> from unyt import unyt_array
>>> data_with_units = unyt_array(data, g)
>>> data_with_units.base is data
True
Any set of units can be used for either of these operations. For example, if you already have an existing array, you could do this to create a new array with the same units:
>>> more_data = [4, 5, 6]*data_with_units.units
>>> more_data
unyt_array([4, 5, 6], 'g')
Working with code that uses astropy.units¶
The unyt library can convert data contained inside of an Astropy
Quantity instance. It can also produce a Quantity from an existing
unyt_array instance. To convert data from
astropy.units to unyt use the unyt_array.from_astropy function:
>>> from astropy.units import km
>>> from unyt import unyt_quantity
>>> unyt_quantity.from_astropy(km)
unyt_quantity(1., 'km')
>>> a = [1, 2, 3]*km
>>> a
<Quantity [1., 2., 3.] km>
>>> unyt_array.from_astropy(a)
unyt_array([1., 2., 3.], 'km')
To convert data to astropy.units use the unyt_array.to_astropy method:
>>> from unyt import g, cm
>>> data = [3, 4, 5]*g/cm**3
>>> data.to_astropy()
<Quantity [3., 4., 5.] g / cm3>
>>> (4*cm).to_astropy()
<Quantity 4. cm>
Working with code that uses Pint¶
The unyt library can also convert data contained in Pint Quantity
instances. To convert data from Pint to unyt, use the unyt_array.from_pint function:
>>> from pint import UnitRegistry
>>> import numpy as np
>>> ureg = UnitRegistry()
>>> a = np.arange(4)
>>> b = ureg.Quantity(a, "erg/cm**3")
>>> b
<Quantity([0 1 2 3], 'erg / centimeter ** 3')>
>>> c = unyt_array.from_pint(b)
>>> c
unyt_array([0, 1, 2, 3], 'erg/cm**3')
And to convert data contained in a unyt_array
instance, use the unyt_array.to_pint
method:
>>> from unyt import cm, s
>>> a = 4*cm**2/s
>>> print(a)
4 cm**2/s
>>> a.to_pint()
<Quantity(4, 'centimeter ** 2 / second')>
>>> b = [1, 2, 3]*cm
>>> b.to_pint()
<Quantity([1 2 3], 'centimeter')>
Reading quantities from text¶
Quantities can also be parsed from strings with the unyt_quantity.from_string function:
>>> from unyt import unyt_quantity
>>> unyt_quantity.from_string("1 cm")
unyt_quantity(1, 'cm')
>>> unyt_quantity.from_string("1e3 Msun")
unyt_quantity(1000., 'Msun')
>>> unyt_quantity.from_string("1e-3 g/cm**3")
unyt_quantity(0.001, 'g/cm**3')
This method is helpful to read data from text files, for instance configuration files. It is intended to be as flexible as possible on the string format, though it requires that the numerical value and the unit name be separated with some kind of whitespace.
User-Defined Units¶
Often it is convenient to define new custom units. This can happen when you need
to make use of a unit that the unyt library does not have a definition
for already. It can also happen when dealing with data that uses a custom unit
system or when writing software that needs to deal with such data in a flexible
way, particularly when the units might change from dataset to dataset. This
comes up often when modeling a physical system since it is often convenient to
rescale data from a physical unit system to an internal “code” unit system in
which the values of the variables under consideration are close to unity. This
approach can help minimize floating point round-off error but is often done for
convenience or to non-dimensionalize the problem under consideration.
The unyt library provides two approaches for dealing with this
problem. For more toy one-off use-cases, we suggest using
unyt.define_unit which allows defining a
new unit name in the global, default unit system that unyt ships with by
default.
This function makes it possible to easily define a new unit that is unknown to
the unyt library:
>>> import unyt as u
>>> ninety_pounds = 90.0*u.lb
>>> one_pound = 1.0*u.lb
>>> u.define_unit("firkin", ninety_pounds)
>>> print((3*u.firkin)/one_pound)
270.0 dimensionless
This is primarily useful for one-off definitions of units that the unyt
library does not already have predefined. For more complex uses cases that need
more flexibility, it is possible to use a custom unit system by ensuring that
the data you are working with makes use of a UnitRegistry customized for your use case, as described
below.
Dealing with data types¶
The unyt library supports creating unyt.unyt_array and unyt.unyt_quantity instances with arbitrary integer or floating point
data types:
>>> import numpy as np
>>> from unyt import km
...
>>> int_data = [1, 2, 3]*km
>>> int_data
unyt_array([1, 2, 3], 'km')
>>> float32_data = np.array([1, 2, 3], dtype='float32')*km
>>> float32_data
unyt_array([1., 2., 3.], dtype=float32, units='km')
The dtype of a unyt_array instance created by multiplying an iterable by
a unit will be the same as passing the iterable to np.array(). You can also
manually specify the dtype by calling np.array() yourself or by using
the unyt_array initializer directly:
>>> np.array([1, 2, 3], dtype='float64')*km
unyt_array([1., 2., 3.], 'km')
Operations that convert an integer array to a new unit will convert the array to
the floating point type with an equivalent size. For example, Calling
in_units on a 32 bit integer array with units of kilometers will return a 32
bit floating point array.
>>> data = np.array([1, 2, 3], dtype='int32')*km
>>> data.in_units('mile')
unyt_array([0.6213712, 1.2427424, 1.8641136], dtype=float32, units='mile')
In-place operations will also mutate the dtype from float to integer in these cases, again in a way that will preserve the byte size of the data.
>>> data.convert_to_units('mile')
>>> data
unyt_array([0.6213712, 1.2427424, 1.8641136], dtype=float32, units='mile')
It is possible that arrays containing large integers (16777217 for 32 bit and 9007199254740993 for 64 bit) will lose precision when converting data to a different unit. In these cases a warning message will be printed.
Integrating unyt Into a Python Library¶
The unyt library began life as the unit system for the yt data
analysis and visualization package, in the form of yt.units. In this role,
unyt was deeply integrated into a larger Python library. Due to these
origins, it is straightforward to build applications that ensure unit
consistency by making use of unyt. Below we discuss a few topics that
most often come up when integrating unyt into a new or existing Python
library.
Unit registries¶
It is also possible to define a custom database of units completely independent
of the global default unit database exposed by the unyt namespace or to
create namespaces in your own package that expose listings of units. In these
cases it becomes important to understand how unyt stores unit metadata in an
internal database, how to add custom entries to the database, how to modify
them, and how to persist custom units.
In practice, the unit metadata for a unit object is contained in an instance of the UnitRegistry class. Every Unit instance contains a reference to a UnitRegistry instance:
>>> from unyt import g
>>> g.registry
<unyt.unit_registry._NonModifiableUnitRegistry ...>
All the unit objects in the unyt namespace make use of the default unit
registry, importable as unyt.unit_registry.default_unit_registry. This
registry object contains all of the real-world physical units that the
unyt library ships with out of the box.
The unit registry itself contains a look-up table that maps from unit names to the metadata necessary to construct a unit. Note that the unit registry only contains metadata for “base” units, and not, for example, SI-prefixed units like centimeter of kilogram, it will instead only contain entries for meter and gram.
Sometimes it is convenient to create a unit registry containing new units that are not available in the default unit registry. A common example would be adding a code_length unit that corresponds to the scaling to from physical lengths to an internal unit system. In practice, this value is arbitrary, but will be fixed for a given problem. Let’s create a unit registry and a custom "code_length" unit to it, and then create a "code_length" unit and a quantity with units of "code_length". For the sake of example, let’s set the value of "code_length" equal to 10 meters.
>>> from unyt import UnitRegistry, Unit
>>> from unyt.dimensions import length
>>> reg = UnitRegistry()
>>> reg.add("code_length", base_value=10.0, dimensions=length,
... tex_repr=r"\rm{Code Length}")
>>> 'code_length' in reg
True
>>> u = Unit('code_length', registry=reg)
>>> data = 3*u
>>> print(data)
3 code_length
As you can see, you can test whether a unit name is in a registry using the
Python in operator.
In an application that depends on unyt, it is often convenient to define
methods or functions to automatically attach the correct unit registry to unit
objects associated with an object. For example, consider a Simulation
class. Let’s give this class two methods named array and quantity to
create new unyt_array and unyt_quantity instances, respectively:
>>> class Simulation:
... def __init__(self, registry):
... self.registry = registry
...
... def array(self, value, units):
... return unyt_array(value, units, registry=self.registry)
...
... def quantity(self, value, units):
... return unyt_quantity(value, units, registry=self.registry)
...
>>> registry = UnitRegistry()
>>> registry.add("code_length", base_value=3.2, dimensions=length)
>>> s = Simulation(registry)
>>> s.array([1, 2, 3], 'code_length')
unyt_array([1, 2, 3], 'code_length')
We can create an array with "code_length" here because s.registry, the UnitRegistry instance associated with our Simulation instance has a "code_length" unit defined.
As for arrays with different units, for operations between arrays created with different unit registries, the result of the operation will use the same unit registry as the leftmost unit. This can sometimes lead to surprising behaviors where data will seem to “forget” about custom units. In this situation it is important to make sure ahead of time that all data are created with units using the same unit registry. If for some reason that is not possible (for example, when comparing data from two different simulations with different internal units), then care must be taken when working with custom units. To avoid these sorts of ambiguities it is best to do work in physical units as much as possible.
When writing tests, it is convenient to use unyt.testing. In particular, assert_allclose_units can be used to check for floating-point equality.
>>> from unyt import assert_allclose_units, m
>>> import numpy as np
>>> actual = [1e-5, 1e-3, 1e-1] * m
>>> desired = actual.to("cm")
>>> assert_allclose_units(actual, desired)
Custom Unit Systems¶
By default unyt uses the SI MKS unit system. However, libraries can
create a unit registry using another unit system to expose that unit system to
their users by creating a unit registry with a custom unit system. For example,
to make CGS units the default unit for all operations, one might use a CGS
UnitRegistry to instancitate the Simulation class like so:
>>> class Simulation:
... def __init__(self, registry):
... self.registry = registry
...
... def array(self, value, units):
... return unyt_array(value, units, registry=self.registry)
...
... def quantity(self, value, units):
... return unyt_quantity(value, units, registry=self.registry)
...
>>> registry = UnitRegistry(unit_system='cgs')
>>> registry.add("code_length", base_value=3.2, dimensions=length)
>>> s_cgs = Simulation(registry)
>>> data = s_cgs.array([1, 2, 3], 'code_length')
>>> data
unyt_array([1, 2, 3], 'code_length')
>>> data.in_base()
unyt_array([320., 640., 960.], 'cm')
Note that the base_value parameter of UnitRegistry.add must be specified in MKS units. All unit
data are stored internally in unyt in MKS units.
You can also use two helper functions provided by unyt,
unyt.unit_systems.add_constants() and
unyt.unit_systems.add_symbols(), to populate a namespace with a set of
predefined unit symbols or physical consants. This namespace could correspond to
the names importable from a module or the names of attributes of an object, or
any other generic dictionary.
One example of doing this would be to make a UnitContainer class that
contains units that are compatible with the Simulation instance we named
s_cgs in the example above:
>>> from unyt.unit_systems import add_symbols
>>> class UnitContainer:
... def __init__(self, registry):
... add_symbols(vars(self), registry)
>>> units = UnitContainer(s_cgs.registry)
>>> units.kilometer
km
>>> units.code_length
code_length
>>> (10.0 * units.kilometer).in_base()
unyt_quantity(1000000., 'cm')
>>> (10.0 * units.kilometer).in_units('code_length')
unyt_quantity(3125., 'code_length')
Note how the result of the call to in_base() comes out in centimeters
because of the the CGS unit system used by the UnitRegistry instance associated with the Simulation.
Writing Data with Units to Disk¶
The unyt library has support for serializing data stored in a
unyt.unyt_array instance to HDF5 files, text
files, and via the Python pickle protocol. We give brief examples below, but first describe how to handle saving units manually as string metadata.
Dealing with units as strings¶
If all you want to do is save data to disk in a physical unit or you are working in a physical unit system, then you only need to save the unit name as a string and treat the array data you are trying to save as a regular NumPy array, as in this example:
>>> import numpy as np
>>> import os
>>> from unyt import cm
...
>>> data = [1, 2, 3]*cm
>>> np.save('my_data_cm.npy', data)
>>> new_data = np.load('my_data_cm.npy')
>>> new_data
array([1, 2, 3])
>>> new_data_with_units = new_data * cm
>>> os.remove('my_data_cm.npy')
Of course in this example using numpy.save we need to hard-code the units because the .npy format doesn’t have a way to store metadata along with the array data. We could have stored metadata in a sidecar file, but this is much more natural with hdf5 via h5py:
>>> import h5py
>>> import os
>>> from unyt import cm, unyt_array
...
>>> data = [1, 2, 3]*cm
...
>>> with h5py.File('my_data.h5', 'a') as f:
... d = f.create_dataset('my_data', data=data)
... f['my_data'].attrs['units'] = str(data.units)
...
>>> with h5py.File('my_data.h5', 'r') as f:
... new_data = f['my_data'][:]
... unit_str = f['my_data'].attrs['units']
...
>>> new_data = unyt_array(new_data, unit_str)
>>> new_data
unyt_array([1, 2, 3], 'cm')
>>> os.remove('my_data.h5')
HDF5 Files¶
The unyt library provides a hook for writing data both to a new HDF5 file and an existing file and then subsequently reading that data back in to restore the array. This works via the unyt_array.write_hdf5 and unyt_array.from_hdf5 methods. The simplest way to use these functions is to write data to a file that does not exist yet:
>>> from unyt import cm, unyt_array
>>> import os
>>> data = [1, 2, 3]*cm
>>> data.write_hdf5('my_data.h5')
...
>>> unyt_array.from_hdf5('my_data.h5')
unyt_array([1, 2, 3], 'cm')
>>> os.remove('my_data.h5')
By default the data will be written to the root group of the HDF5 file in a dataset named 'array_data'. You can also specify that you would like
the data to be saved in a particular group or dataset in the file:
>>> data.write_hdf5('my_data.h5', dataset_name='my_special_data',
... group_name='my_special_group')
>>> unyt_array.from_hdf5('my_data.h5', dataset_name='my_special_data',
... group_name='my_special_group')
unyt_array([1, 2, 3], 'cm')
>>> os.remove('my_data.h5')
You can even write to files and groups that already exist:
>>> with h5py.File('my_data.h5', 'w') as f:
... g = f.create_group('my_custom_group')
...
>>> data.write_hdf5('my_data.h5', group_name='my_custom_group')
...
>>> with h5py.File('my_data.h5') as f:
... print(f['my_custom_group/array_data'][:])
[1 2 3]
>>> os.remove('my_data.h5')
If the dataset that you would like to write to already exists, unyt
will clobber that dataset.
Note that with this method of saving data to HDF5 files, the
unyt.UnitRegistry instance associated
with the units of the data will be saved in the HDF5 file. This means that if
you create custom units and save a unit to disk, you will be able to convert
data to those custom units even if you are dealing with those units later after
restoring the data from disk. Here is a short example illustrating this:
>>> import os
>>> from unyt import UnitRegistry
>>> reg = UnitRegistry()
>>> reg.add("code_length", base_value=10.0, dimensions=length,
... tex_repr=r"\rm{Code Length}")
>>> u = Unit('cm', registry=reg)
>>> data = [1., 2., 3.]*u
>>> data.write_hdf5('my_code_data.h5')
>>> read_data = data.from_hdf5('my_code_data.h5')
>>> read_data
unyt_array([1., 2., 3.], 'cm')
>>> read_data.to('code_length')
unyt_array([0.001, 0.002, 0.003], 'code_length')
>>> os.remove('my_code_data.h5')
Text Files¶
The unyt library also has wrappers around numpy.savetxt and numpy.loadtxt for saving data as an ASCII table. For example:
>>> import unyt as u
>>> import os
>>> data = [[1, 2, 3]*u.cm, [4, 5, 6]*u.kg]
>>> u.savetxt('my_data.txt', data)
>>> with open('my_data.txt') as f:
... print("".join(f.readlines()))
# Units
# cm kg
1.000000000000000000e+00 4.000000000000000000e+00
2.000000000000000000e+00 5.000000000000000000e+00
3.000000000000000000e+00 6.000000000000000000e+00
>>> os.remove('my_data.txt')
Pickles¶
Note
Pickle files are great for serializing data to disk or over a network for internal usage by a package. They are ill-suited for long-term data storage or for communicating data between different Python installations. If you want to use pickle files for data storage, consider using a format designed for long-term data storage, like HDF5.
Both unyt.unyt_array and unyt.Unit instances can be saved using the pickle protocol:
>>> from unyt import kg
>>> import pickle
>>> import numpy as np
...
>>> assert kg == pickle.loads(pickle.dumps(kg))
>>> data = [1, 2, 3]*kg
>>> reloaded_data = pickle.loads(pickle.dumps(data))
>>> assert np.array_equal(data.value, reloaded_data.value)
>>> assert data.units == reloaded_data.units
As for HDF5 data, the unit registry associated with the unit object is saved to the pickle. If you have custom units defined, the reloaded data will know about your custom unit and be able to convert data to and from the custom unit.
Handling errors from unyt¶
unyt sometimes raises exceptions with unique exception types, e.g., to signal
invalid operations, like summation of quantities with different dimensions.
It is possible to catch any exceptions from unyt as
>>> from unyt import cm, s
>>> from unyt.exceptions import UnytError
>>> a = 1 * cm
>>> b = 1 / s
>>> try:
... a + b
... except UnytError:
... pass
However, it is in general advised to only catch specific exceptions types that
are known-possible outcomes. All custom exceptions types live in the
unyt.exceptions module and may be imported from there.
Performance Considerations¶
Tracking units in an application will inevitably add overhead. Judging where overhead is important or not depends on what real-world workflows look like. Ultimately, profiling code is the best way to find out whether handling units is a performance bottleneck. Optimally handling units will be amortized over the cost of an operation. While this is true for large arrays (bigger than about one million elements), this is not true for small arrays that contain only a few elements.
In addition, it is sometimes easy to write code that needlessly checks unit consistency when we know ahead of time that data are already in the correct units. Often we can get away with only checking unit consistency once and then stripping units after that.
A good rule of thumb is that units should be checked on input, stripped off of data during a calculation, and then re-applied when returning data from a function. In other words, apply or check units at interfaces, but during an internal calculation it is often worth stripping units, especially if the calculation involves many operations on arrays with only a few elements.
unyt_array.name attribute¶
The unyt_array has a name attribute for use in structured-data applications or similar applications that require labeled data. For example, Numpy has record arrays and when constructed as shown below, it is possible to retain the units while taking advantage of the labeled record fields.
>>> import numpy as np
>>> from unyt import unyt_array
>>> x = unyt_array([0, 1, 2], "s", name="time")
>>> y = unyt_array([3, 4, 5], "m", name="distance")
>>> data = (x, y)
>>> dt = [(a.name, "O") for a in data]
>>> data_points = np.array(list(zip(*data)), dtype=dt).view(np.recarray)
>>> data_points[0].time
unyt_quantity(0, 's')
>>> data_points[0].distance
unyt_quantity(3, 'm')
Note
The name attribute does not propagate through mathematical operations. Other operations such as indexing, copying, and unit conversion, will preserve the name attribute where the semantic meaning of the quantity remains the same.
Plotting with Matplotlib¶
Note
This is an experimental feature. Please report issues.
This feature works in Matplotlib versions 2.2.4 and above
Matplotlib is not a dependency of Unyt
Matplotlib is Unyt aware. After enabling support in unyt using the
unyt.matplotlib_support context
manager, Matplotlib will label the x and y axes with the units.
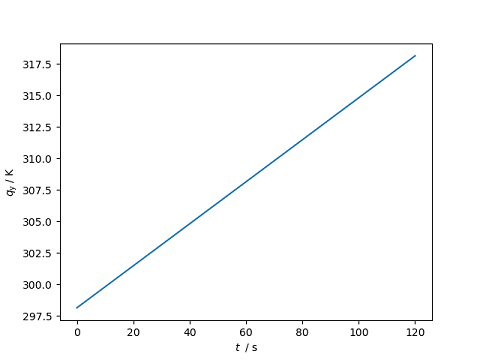
>>> import matplotlib.pyplot as plt
>>> from unyt import matplotlib_support, s, K
>>> x = [0.0, 60.0, 120.0]*s
>>> y = [298.15, 308.15, 318.15]*K
>>> with matplotlib_support:
... plt.plot(x, y)
... plt.show()
[<matplotlib.lines.Line2D object at ...>]
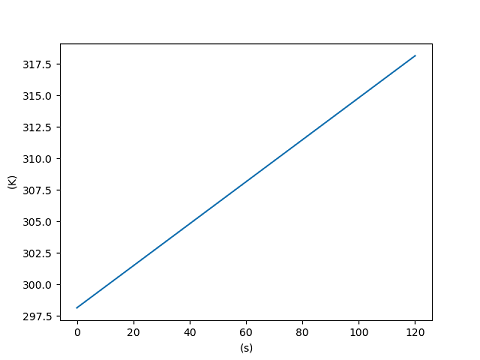
You can change the plotted units without affecting the original data.
>>> with matplotlib_support:
... plt.plot(x, y, xunits="min", yunits=("J", "thermal"))
... plt.show()
[<matplotlib.lines.Line2D object at ...>]
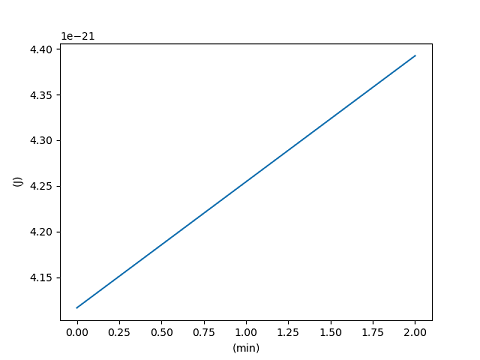
It is also possible to set the label style; the choices "()", "[]" and
"/" are supported.
>>> matplotlib_support.label_style = "[]"
>>> with matplotlib_support:
... plt.plot(x, y)
... plt.show()
[<matplotlib.lines.Line2D object at ...>]
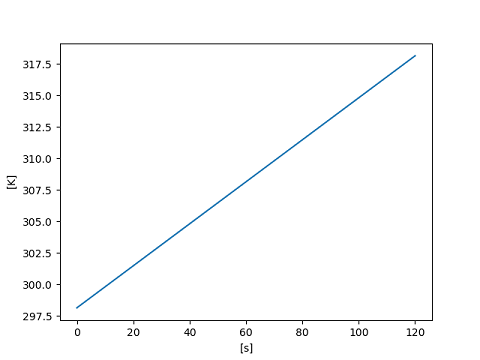
The axis label will include the unyt_array.name attribute if set.
>>> x.name = "Time"
>>> y.name = "Temperature"
>>> with matplotlib_support:
... plt.plot(x, y)
... plt.show()
[<matplotlib.lines.Line2D object at ...>]
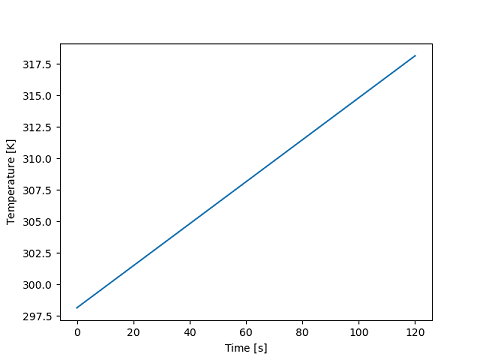
With label_style set to “/”, the axis label conforms to the SI standard where the axis label is a mathematical expression rather than a caption. In this case, set the unyt_array.name attribute to the latex expression for the physical quantity symbol.
>>> x.name = "$t$"
>>> y.name = ""
>>> matplotlib_support.label_style = "/"
>>> with matplotlib_support:
... plt.plot(x, y)
... plt.show()
[<matplotlib.lines.Line2D object at ...>]
There are three ways to use the context manager:
As a conventional context manager in a
withstatement as shown aboveAs a feature toggle in an interactive session:
>>> import matplotlib.pyplot as plt
>>> from unyt import s, K, matplotlib_support
>>> matplotlib_support.enable()
>>> plt.plot([0, 1, 2]*s, [3, 4, 5]*K)
[<matplotlib.lines.Line2D object at ...>]
>>> plt.show()
>>> matplotlib_support.disable()
As an enable for a complete session:
>>> import unyt
>>> unyt.matplotlib_support()
>>> import matplotlib.pyplot as plt
Working with Dask arrays¶
unyt provides the ability to wrap dask arrays with unyt
behavior. The main access point is the unyt.dask_array.unyt_from_dask
function, which allows you to build a unyt_dask_array from a plain dask array
analogous to the creation of a unyt_array from a plain numpy.ndarray:
>>> from unyt import dask_array as uda
>>> import dask.array as da
>>> x = da.arange(10000, chunks=(1000,))
>>> x_da = uda.unyt_from_dask(x, 'm')
Methods that hang off of a unyt_dask_array object and operations on
unyt_dask_array objects will generally preserve units:
>>> x_da.sum().compute()
unyt_quantity(49995000, 'm')
>>> (x_da[:5000] * x_da[5000:]).compute()[:5]
unyt_array([ 0, 5001, 10004, 15009, 20016], 'm**2')
One important caveat is that using Dask array functions may strip units:
>>> da.sum(x_da).compute()
49995000
For simple reductions, you can use the reduce_with_units function:
>>> result = uda.reduce_with_units(da.sum, x_da)
>>> result.compute()
unyt_quantity(49995000, 'm')
But more complex operations may require more careful management of units. Note
that reduce_with_units will accept any of the positional or keyword
arguments for the array function:
>>> import numpy as np
>>> x = da.ones((10000, 3), chunks=(1000, 1000))
>>> x[:,0] = np.nan
>>> x_da = uda.unyt_from_dask(x, 'm')
>>> result = uda.reduce_with_units(da.nansum, x_da, axis=1)
>>> result.compute()[:5]
unyt_array([2., 2., 2., 2., 2.], 'm')
As a final note: the initial Dask array provided to dask_array.unyt_from_dask can be
constructed in any of the usual ways of constructing Dask arrays – from NumPy-like
array instantiation as in the above examples to reading from file or delayed operations.
For more on creating arrays, check out the Dask documentation.